Higher-order functional domains in the human ENCODE regions
This webpage is intended as a supplement to the manuscript, "Higher-order functional domains in the human ENCODE regions," by R.E. Thurman, N. Day, W.S. Noble, and J.A. Stamatoyannopoulos, Genome Research, 2007, 17, 991-994. Below find data, links to data, and analysis results referenced in that paper. There are also some results here included in the ENCODE Nature paper, "Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project", Nature, 2007, 447, 799-816 (see 6-track segmentation below).As part of the ENCODE project we have access to a number of datatypes, representing a variety of functional variables (e.g., DNaseI sensitivity, transcription, histone modifications, conservation, etc.) sampled in a nearly continuous fashion across the genome. We would like to understand to what degree such data reveal coherent higher-order features that may in turn illuminate the underlying functional architecture of the genome. To address this, we aim to develop approaches based on wavelet analysis for the discovery of "domain-level" behavior in fine scale data, and for correlating these apparently disparate functional data types.
The rough outline of methods used in the paper are as follows:
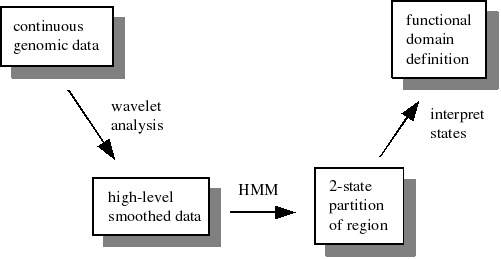
(postscript version of this diagram)
- Use wavelets and HMMs to define functional domains based on one or
more continuous functional datasets from the ENCODE project. Those
datasets, and their public availability, are listed below. Datasets
available from the UCSC Genome
Browser were downloaded under the ENCODE section using hg17 (May,
2004 assembly) coordinates.
- Histone modifications
- H3ac, H4ac, H3K4me1, H3K4me2, H3K4me3 available from the Sanger institute, UCSC Genome Browser tracks encodeSangerChipH3acHela, encodeSangerChipH4acHela, encodeSangerChipH3K4me1Hela, encodeSangerChipH3K4me2Hela, and encodeSangerChipH3K4me3Hela.
- H3K27me3 available from the University of San Diego, UCSC Genome Browser track encodeUcsdChipH3K27me3.
- RNA expression levels from Affymetrix, UCSC Genome Browser track encodeAffyRnaHeLaSignal
- DNA replication timing from the University of Virgina, UCSC Genome Browser track encodeUvaDnaRepTr50
- density of conserved non-coding sequence elements, from the ENCODE
MSA (multi-species alignment) analysis subgroup. This data to appear
on the UCSC Genome Browser. Links to the actual data used for the
paper are here:
- MSA "moderate" non-exonic raw data (coordinates of non-exonic conserved elements defined using the MSA group's "moderate" criteria).
- Density of above elements in 3kb windows stepping at 1kb intervals
- "Validate" these domains in a variety of ways. For instance,
exhibit statistically significant differences between HMM states in
- percent coding sequence
- # tx start sites
- # CNS
- MSA moderate non-exonic (number of these bases in a sliding window)
- % CpG content
Software
- For wavelet smoothing using the maximal-overlap discrete wavelet transform (MODWT), we use R package waveslim.
- For HMM segmentation we use our own software package, HMMSeg (manuscript "Unsupervised segmentation of continuous genomic data," currently in review).
- Utility routines in R for data pre-processing (interpolation using adaptive loess fitting is included in R function UCSC2bed in file bed.R.
Annotation results
Segmentations
Single-track segmentations
- Sanger Chip (HeLa), 64kb wavelet smooth
- Affy RNA Signal, 51.2kb wavelet smooth
- UCSD Chip, H3K27me3, 64kb wavelet smooth
- UVa DNA Replication, TR50
- MSA non-exonic, 3kb sliding window, 64kb wavelet smooth
Multiple-track segmentations
- 3 track: H3ac, H3K27me3, and TR50 (HeLa), wavelet-smoothed as above
- 4 track: Affy RNA, H3ac, H3K27me3, and TR50 (HeLa), wavelet-smoothed as above
- 5 track: Affy RNA, H3ac, H3K27me3, MSA non-exonic, and TR50 (HeLa), wavelet-smoothed as above
- 6 track: Affy RNA, H3ac, H3K27me3, TR50, merged HS density, RFBR density (HeLa) -- for ENCODE Nature paper
Segmentation comparisons
Concordance figures are the percentage of bases whose state assignments agree in both segmentations.Single-track comparisons
Sanger histone modifications vs. themselves
Tissue differences: HeLa vs. GM
- Sanger H3K4me2, 64kb smooth 76% concordance
- Sanger H3K4me1, 64kb smooth 69% concordance
- Sanger H3K4me3, 64kb smooth 78% concordance
- Sanger H3ac, 64kb smooth 72% concordance
- Sanger H4ac, 64kb smooth 76% concordance
- Affy RNA Signal, 51.2kb smooth 81% concordance
Wavelet scale differences
- SangerH3K4me2, GM, 2kb smooth vs. 64kb smooth 70% concordance
- SangerH3K4me2, segment size as a function of scale (postscript file)
- Segment boundary scale sensitivity, based on 4-track (Affy zero-threshold, H3ac, H3K27me3, and TR50) segmentations at 32kb, 64kb, and 128kb scales.
Affy/H3ac/H3K27me3/TR50 wavelet 4-track vs. each of its component single-track results
- Sanger H3ac 64kb (89% concordance)
- Affy RNA, 51.2kb (80% concordance)
- UCSD Chip, H3K27me3 64kb (62% concordance)
- TR50 (76% concordance).
Affy/H3ac/H3K27me3/MSA non-exonic/TR50 wavelet 5-track vs. each of its component single-track results
- Sanger H3ac 64kb (90% concordance).
- Affy RNA, 51.2kb (79% concordance).
- UCSD Chip, H3K27me3 64kb (62% concordance).
- TR50 (76% concordance).
- MSA non-exonic (62% concordance)
3-track vs. 4-track
- H3ac/H3K27me3/Tr50 vs Affy/H3ac/H3K27me3/MSA/Tr50, 78% concordance
4-track vs. 4-track
- Affy/H3ac/H3K27me3/Tr50 vs. H3ac/H3K27me3/MSA/Tr50, 78% concordance
4-track vs. 5-track
| Affy RNA Signal | UCSD Chip H3K27me3 | MSA non-exonic | Uva TR50 | |
| Sanger H3K4me1 | 67% concordance | 62% concordance | 56% concordance | 74% concordance |
| Sanger H3K4me2 | 72% concordance | 55% concordance | 58% concordance | 69% concordance |
| Sanger H3K4me3 | 68% concordance | 52% concordance | 53% concordance | 60% concordance |
| Sanger H3ac | 75% concordance | 59% concordance | 63% concordance | 70% concordance |
| Sanger H4ac | 74% concordance | 63% concordance | 61% concordance | 75% concordance |
| Affy RNA Signal | 49% concordance | 57% concordance | 61% concordance | |
| UCSD Chip H3K27me3 | 54% concordance | 70% concordance | ||
| MSA non-exonic | 57% concordance |
| H3ac | H4ac | H3K4me1 | H3K4me2 | H3K4me3 | |
| H3ac | |||||
| H4ac | 85% concordance | ||||
| H3K4me1 | 73% concordance | 80% concordance | |||
| H3K4me2 | 86% concordance | 80% concordance | 77% concordance | ||
| H3K4me3 | 80% concordance | 73% concordance | 68% concordance | 87% concordance |
High-confidence segments
- 4 track: Affy (zero threshold), H3ac, H3K27me3, TR50 (wavelet)
- 5 track: Affy (zero threshold), H3ac, H3K27me3, MSA non-exonic, TR50 (wavelet)
- Affy, H3ac, H3K27me3, TR50 - single-track segmentations only