A statistical framework for genomic data fusion
Gert R. G. Lanckriet, Tijl De Bie, Nello Cristianini, Michael I. Jordan and William Stafford Noble.Supplement (PDF)
Data and kernel matrices
In the following table, the data and kernel matrices are provided as tab-delimited text files, with string IDs in the first row and column. Warning: Some of the kernel matrices are quite large and may cause your browser to hang if you try to load them directly. You can save them to your local disk by right clicking on the link.The links marked "PNG" are heat map representations of the corresponding matrices, generated using matrix2png. For the data matrices, the color range is selected to span the middle 98% of the values in the matrix.
| KB | BLAST | Data (PNG) | Kernel (PNG) | CN |
| KSW | Smith-Waterman | Data (PNG) | Kernel (PNG) | CN |
| KPfam | Pfam HMM | Data (PNG) | Kernel (PNG) | N |
| KFFT | Hydrophobicity FFT | Data (PNG) | Kernel (PNG) | CN |
| KLI | Linear interactions | Data (PNG) | Kernel (PNG) | CN |
| KD | Diffusion kernel | Data (PNG) | Kernel (PNG) | N |
| KE | Gene expression | Data (PNG) | Kernel (PNG) | CN |
| KRND | Random | Data (PNG) | Kernel (PNG) | N |
In order to evaluate the kernel matrices for the prediction problems further on, the relevant submatrices (corresponding to the genes with known classification) need to be selected, and then either normalized (N) or centered and then normalized (CN), as indicated in the last column of the table. When predicting unannotated genes or proteins, the full kernel matrices are needed and should accordingly be centered or not and then normalized.
Matlab code to center a kernel matrix can be found here and Matlab code for normalization here.
Ribosomal proteins
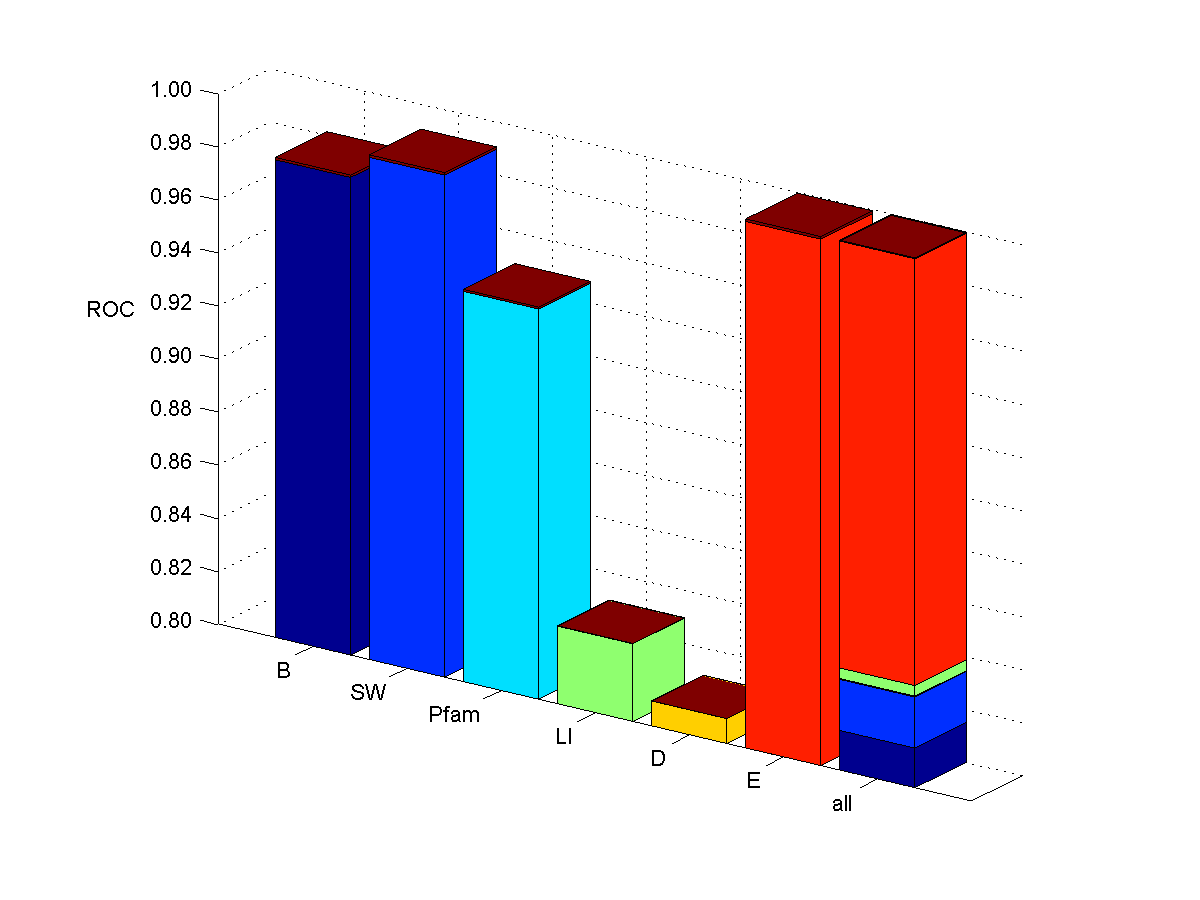
- List of 1150 yeast ORF IDs corresponding to genes known to participate in complexes
- The corresponding FASTA file of sequences.
- Annotations of each gene as ribosomal (1), non-ribosomal (-1), or unknown (0).
Membrane proteins
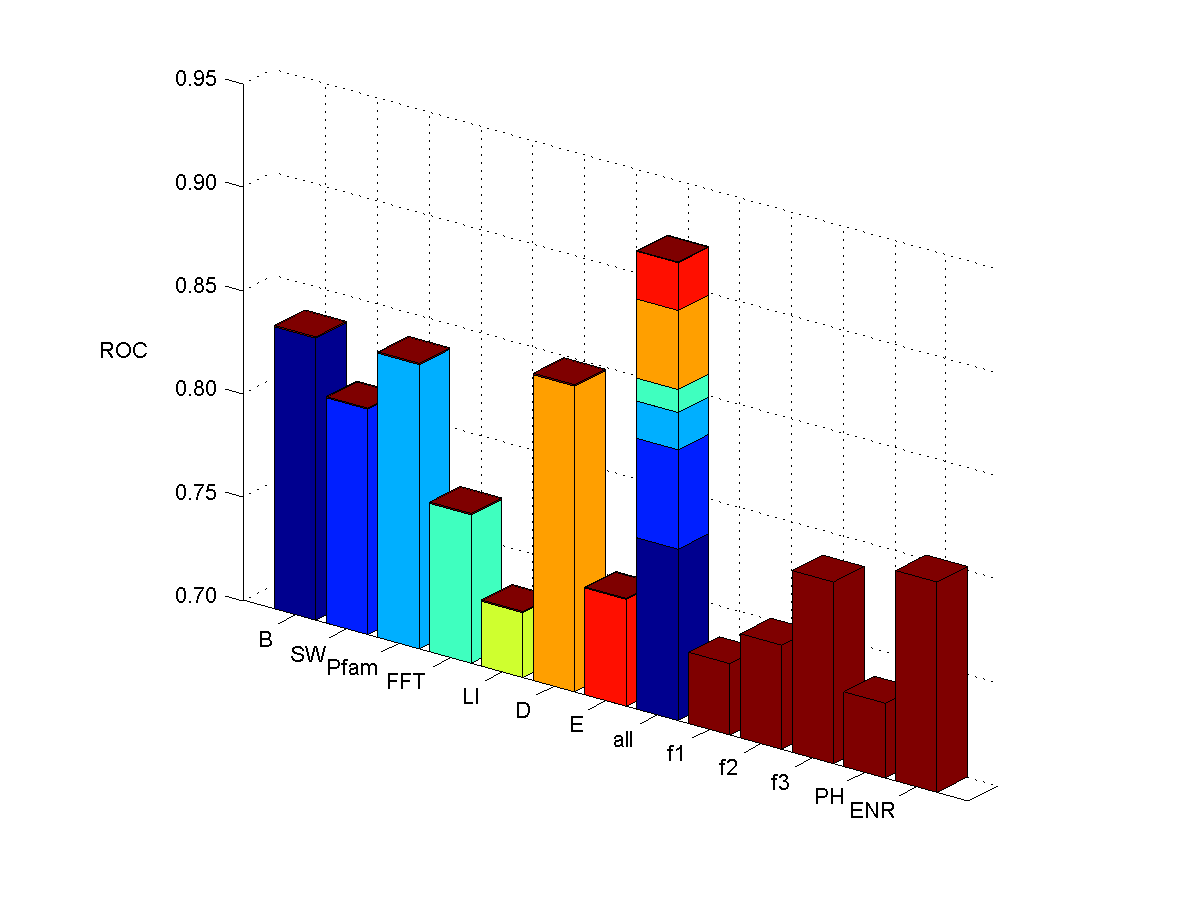
- List of 2318 yeast ORF IDs corresponding to genes with known locations.
- The corresponding FASTA file of sequences.
- Annotations of each gene as membrane (1), non-membrane (-1) or unknown (0).

An example membrane protein. pSR II is a photoreceptor in the plasma membrane of archaeon Natronobacteriumpharaonis and functions as a sensor for phototactic avoidance.