Annotation results
SangerChip data, H3K4me3, HeLa, 64kb wavelet smooth data, 2-state segmentation
|
Gene overlapBed files below contain one line for each gene whose Tx start sits inside the given state. Columns are chr, start, stop, gene ID, fraction of gene sitting inside the segment, strand, and segment ID
|
Overlap with various elements
Total number of bases overlapping various elements in each state. Numbers in parentheses are the fractions of the element in that state. Second set of columns gives enrichment over expected (as a fraction). Enrichment p-values, computed by permuting state labels on the fixed segmentation, are also included. The third set of columns gives fraction of each state's segments that overlap the given element.Element st0 st1 st0 enr st0 enr p st1 enr st1 enr p st0 (frac) st1 (frac) ------------------------- -------------------- -------------------- --------- ---------- --------- ---------- ---------- ---------- Gencode_Known_Exon_Ovr 668204 (0.404) 984439 (0.596) -0.411 0 0.899 0 0.404 0.596 Gencode_Putative_Exon_Ovr 71779 (0.543) 60323 (0.457) -0.208 0.0698 0.455 0.0065 0.543 0.457 Gencode_Pseudo_Exon_Ovr 91426 (0.672) 44606 (0.328) -0.021 0.4415 0.045 0.4009 0.672 0.328 Gencode_Intron_Ovr 11373203 (0.675) 5483194 (0.325) -0.017 0.355 0.037 0.2591 0.675 0.325 Gencode_Coding_Exon_Ovr 373708 (0.463) 433678 (0.537) -0.326 0 0.712 0 0.463 0.537 Gencode_5P.UTR_Ovr 216735 (0.340) 420017 (0.660) -0.504 0 1.102 0 0.340 0.660 Gencode_3P.UTR_Ovr 388721 (0.423) 530533 (0.577) -0.384 0 0.839 0 0.423 0.577 Gencode_1stExon_Ovr 299033 (0.378) 492153 (0.622) -0.449 0 0.983 0 0.378 0.622 KnownGene_Intron_Ovr 9136143 (0.675) 4391651 (0.325) -0.016 0.3949 0.035 0.3279 0.675 0.325 KnownGene_Exon_Ovr 487964 (0.431) 643867 (0.569) -0.372 0 0.813 0 0.431 0.569 KnownGene_5P.UTR_Ovr 54235 (0.379) 88907 (0.621) -0.448 0 0.980 0 0.379 0.621 KnownGene_3P.UTR_Ovr 196828 (0.450) 240844 (0.550) -0.345 0 0.754 0 0.450 0.550 EST_Ovr 12598299 (0.647) 6865709 (0.353) -0.057 0.0623 0.124 0.0042 0.647 0.353 Gene.Unit.Ovr 13653824 (0.660) 7030351 (0.340) -0.038 0.1818 0.083 0.0562 0.660 0.340 CpG.UCSC_Isl_Ovr 140752 (0.356) 254222 (0.644) -0.481 0 1.051 0 0.356 0.644 CpG.AL_Isl_Ovr 438149 (0.417) 613425 (0.583) -0.393 0.0008 0.859 0 0.417 0.583 RptMskr_Ovr 9019200 (0.672) 4399980 (0.328) -0.021 0.0891 0.045 0.014 0.672 0.328 RptMskr_DNA_Ovr 586340 (0.713) 236052 (0.287) 0.039 0.167 -0.085 0.0432 0.713 0.287 RptMskr_LINE_Ovr 3922875 (0.743) 1357347 (0.257) 0.083 0.0081 -0.181 0.0002 0.743 0.257 RptMskr_LINE.L1_Ovr 3132871 (0.751) 1038277 (0.249) 0.094 0.0144 -0.207 0.0003 0.751 0.249 RptMskr_LINE.L2_Ovr 710328 (0.707) 294766 (0.293) 0.030 0.2542 -0.065 0.129 0.707 0.293 RptMskr_LTR_Ovr 1533513 (0.732) 560799 (0.268) 0.067 0.0542 -0.147 0.0026 0.732 0.268 RptMskr_SINE.ALU_Ovr 1983513 (0.518) 1842799 (0.482) -0.245 0 0.535 0 0.518 0.482 RptMskr_SINE_Ovr 2630123 (0.557) 2094541 (0.443) -0.189 0.0002 0.413 0 0.557 0.443 SimpRep_Ovr 515948 (0.687) 234980 (0.313) 0.001 0.5028 -0.003 0.4989 0.687 0.313 Uva_DnaRep.Early_Ovr 3206987 (0.468) 3647745 (0.532) -0.318 0.0014 0.696 0 0.468 0.532 Uva_DnaRep.Mid_Ovr 5480799 (0.708) 2258262 (0.292) 0.032 0.3782 -0.070 0.2866 0.708 0.292 Uva_DnaRep.Late_Ovr 6801827 (0.853) 1169580 (0.147) 0.243 0.0062 -0.532 0 0.853 0.147 Uva_DnaRep.Pans_Ovr 3776971 (0.720) 1466521 (0.280) 0.050 0.3724 -0.109 0.278 0.720 0.280 MSA_strict_Ovr 425672 (0.594) 290656 (0.406) -0.134 0.0563 0.293 0.0012 0.594 0.406 MSA_moderate_Ovr 896738 (0.607) 579890 (0.393) -0.115 0.0371 0.252 0.0004 0.607 0.393 MSA_loose_Ovr 2252707 (0.639) 1275273 (0.361) -0.070 0.0471 0.152 0.0007 0.639 0.361 MSA_loose.NonExon_Ovr 1986059 (0.686) 908901 (0.314) -0.000 0.4979 0.001 0.4963 0.686 0.314 MSA_non.coding_Ovr 715210 (0.707) 296450 (0.293) 0.030 0.384 -0.066 0.294 0.707 0.293 MSA_non.exonic_Ovr 668251 (0.743) 231645 (0.257) 0.082 0.2032 -0.180 0.062 0.743 0.257
Number of overlapping elements
Gives the number of elements of a given category that overlap each state.Element st0 st1 st0 enr st0 enr p st1 enr st1 enr p ------------------------------ -------------------- -------------------- --------- ---------- ---------- ---------- Num_Gencode_TxStarts 1012 (0.357) 1821 (0.643) -0.479 0 1.049 0 Num_Gencode_TxStops 1146 (0.404) 1691 (0.596) -0.411 0 0.900 0 Num_KnownGene_TxStarts 309 (0.379) 507 (0.621) -0.448 0 0.980 0 Num_KnownGene_TxStops 368 (0.448) 454 (0.552) -0.348 0 0.760 0 Num_Ensembl_TxStarts 273 (0.388) 431 (0.612) -0.435 0 0.951 0 Num_Ensembl_TxStops 314 (0.441) 398 (0.559) -0.357 0 0.782 0 Num_mRNA_TxStarts 1449 (0.300) 3386 (0.700) -0.563 0 1.232 0 Num_mRNA_TxStops 1603 (0.334) 3201 (0.666) -0.514 0 1.124 0 Num_Spliced.EST_TxStarts 29806 (0.215) 108569 (0.785) -0.686 0.0008 1.501 0 Num_Spliced.EST_TxStops 25086 (0.271) 67613 (0.729) -0.606 0 1.325 0 Num_CpG_Islands 192 (0.376) 319 (0.624) -0.452 0 0.990 0 Num_SNPs 73908 (0.707) 30595 (0.293) 0.031 0.185 -0.067 0.0489 Num_Nhgri_Chip.GM 707 (0.523) 644 (0.477) -0.237 0 0.519 0 Num_UWDHS.CAP.GM 1390 (0.503) 1375 (0.497) -0.267 0.0001 0.585 0 Num_UWDHS.QCP.GM 366 (0.662) 187 (0.338) -0.036 0.4228 0.078 0.3635 Num_UWDHS.QCP.HeLa 469 (0.668) 233 (0.332) -0.026 0.4375 0.058 0.3936 Num_UVaDnaRep.OriginsPred 177 (0.660) 91 (0.340) -0.038 0.2386 0.082 0.0988 Num_ChipCenters.H3K4me1.GM 1026 (0.604) 672 (0.396) -0.120 0.0002 0.261 0 Num_ChipCenters.H3K4me1.HeLa 527 (0.497) 534 (0.503) -0.276 0 0.604 0 Num_ChipCenters.H3K4me2.GM 802 (0.568) 611 (0.432) -0.173 0 0.378 0 Num_ChipCenters.H3K4me2.HeLa 206 (0.360) 366 (0.640) -0.475 0 1.039 0 Num_ChipCenters.H3K4me3.GM 593 (0.531) 523 (0.469) -0.226 0 0.494 0 Num_ChipCenters.H3K4me3.HeLa 198 (0.346) 374 (0.654) -0.496 0 1.084 0 Num_ChipCenters.H3ac.GM 407 (0.472) 456 (0.528) -0.313 0 0.684 0 Num_ChipCenters.H3ac.HeLa 149 (0.273) 396 (0.727) -0.602 0 1.316 0 Num_ChipCenters.H4ac.GM 701 (0.568) 534 (0.432) -0.173 0.0015 0.378 0 Num_ChipCenters.H4ac.HeLa 287 (0.438) 369 (0.562) -0.362 0 0.793 0 Num_MSA_strict 18147 (0.593) 12463 (0.407) -0.136 0.0177 0.298 0 Num_MSA_moderate 23291 (0.634) 13457 (0.366) -0.076 0.0485 0.167 0.0007 Num_MSA_loose 66526 (0.664) 33724 (0.336) -0.033 0.1383 0.072 0.0224 Num_MSA_loose.NonExon 65156 (0.659) 33722 (0.341) -0.040 0.1127 0.087 0.0117 Num_MSA_non.coding 22221 (0.655) 11692 (0.345) -0.045 0.1871 0.099 0.0479 Num_MSA_non.exonic 21116 (0.677) 10075 (0.323) -0.013 0.404 0.030 0.3264
Enrichment plots
Plot of enrichment values in each state for various elements.
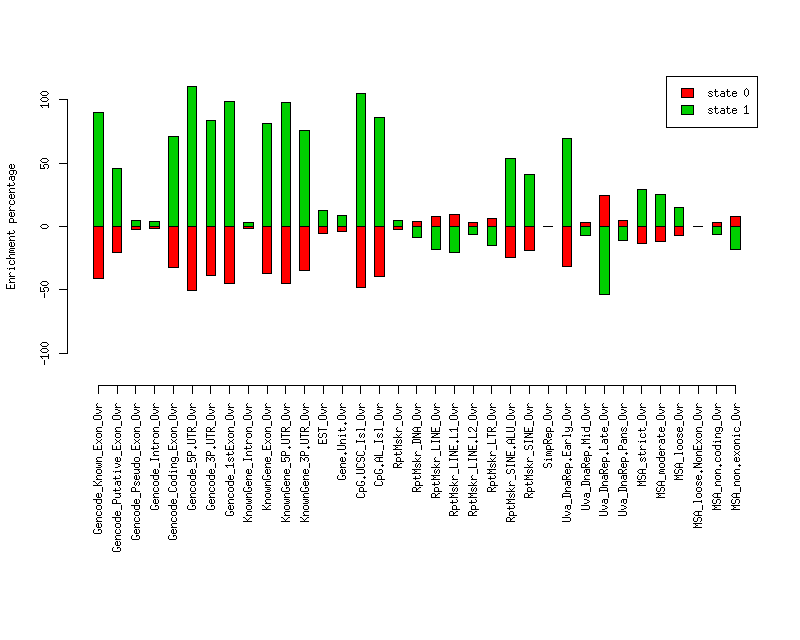
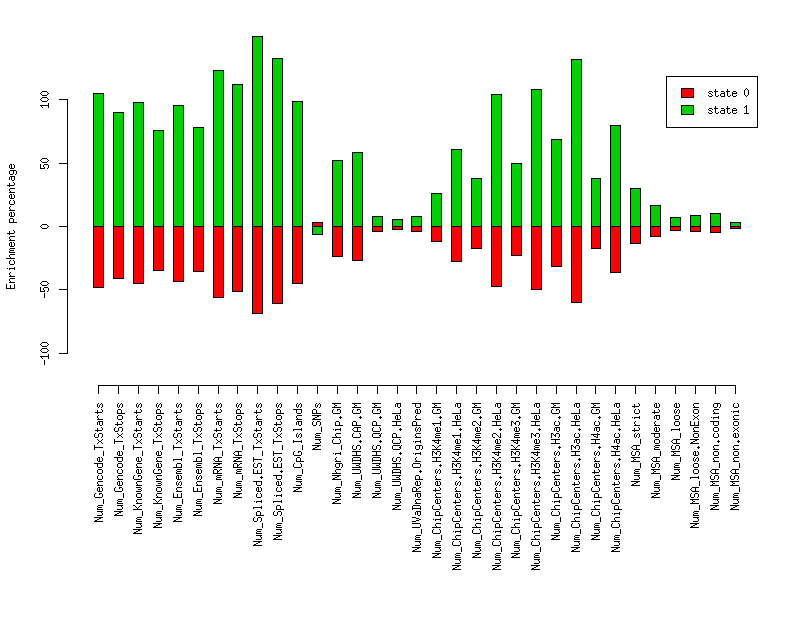
Distances of elements to segment boundaries
Average and median absolute value over all boundaries.
Statistic avg(|d|) med(|d|) ------------------------- ------------ ------------ Gencode_5P_Dist_Known 1791243.56 17839.0 Gencode_3P_Dist_Known 1795727.86 14739.0 Gencode_5P_Dist_Putative 3394362.39 86514.0 Gencode_3P_Dist_Putative 3392627.51 84243.0 Gencode_5P_Dist_Pseudo 1620922.19 79246.0 Gencode_3P_Dist_Pseudo 1621216.66 78614.0 Gencode_TxStop_Dist 791602.52 9448.0 Gencode_TxStart_Dist 792557.76 11025.0 Gencode_5P.UTR_Dist 1809338.08 16146.0 Gencode_1stExon_Dist 793189.19 11067.0 KnownGene_3P_Dist 91468.33 31326.0 KnownGene_5P_Dist 77429.83 29608.0 Ensemble_5P_Dist 79927.53 27412.0 Refseq_5P_Dist 85417.60 34088.0 mRNA_5P_Dist 47821.36 12961.0 mRNA_3P_Dist 48623.81 14746.0 Spliced.EST_5P_Dist 15154.41 3350.0 Spliced.EST_3P_Dist 14518.63 2674.0 CpG.UCSC_Isl_Dist 105279.63 38141.0 CpG.AL_Isl_Dist 645576.29 22709.0 Nhgri_Chip.GM_Dist 618416.07 14163.0 UWDHS.CAP.GM_Dist 640497.79 13348.0 UWDHS.QCP.GM_Dist 19513305.62 58544.0 UWDHS.QCP.HeLa_Dist 19504457.01 53374.0 UVaDnaRep.OriginsPred_Dist 107818.24 38309.0 Sanger_ChipCenters.H3K4me1.GM_Dist 21108.00 9557.0 Sanger_ChipCenters.H3K4me1.HeLa_Dist 819742.17 16371.0 Sanger_ChipCenters.H3K4me2.GM_Dist 26538.67 9863.0 Sanger_ChipCenters.H3K4me2.HeLa_Dist 2644070.95 35254.5 Sanger_ChipCenters.H3K4me3.GM_Dist 39350.97 17257.0 Sanger_ChipCenters.H3K4me3.HeLa_Dist 3335113.33 33901.0 Sanger_ChipCenters.H3ac.GM_Dist 59634.81 24970.0 Sanger_ChipCenters.H3ac.HeLa_Dist 1596835.33 38641.5 Sanger_ChipCenters.H4ac.GM_Dist 48884.79 18707.0 Sanger_ChipCenters.H4ac.HeLa_Dist 1068868.54 38075.0 Uva_DnaRep.Early_Dist 8133206.69 81764.0 Uva_DnaRep.Mid_Dist 3981996.02 44772.5 Uva_DnaRep.Late_Dist 1133565.97 92018.0 Uva_DnaRep.PanS_Dist 4225607.75 159060.0 MSA_non.exonic_Dist 7036.94 2329.0